Physiological Proteomics & Bioinformatics“ (Prof. K. Riedel, Dr. J. Pané-Farré)
The Riedel group focus on the following research topics:
- elucidation of the molecular basis of infections by opportunistic pathogens such as Pseudomonas aeruginosa, Burkholderia sp., Clostridium difficile, Staphylococcus aureus by in vitro and in vivo proteome analyses
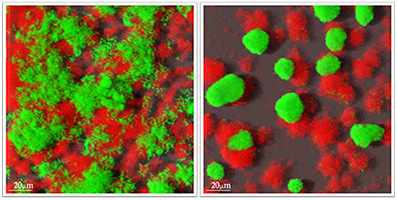
- metaproteomics and microscopic analyses to unravel structure and functions of pathogenic mono- and mixed-species biofilms
- environmental proteomics to study microbial communities in terrestric and aquatic habitats, i.e. leaf litter, soil, lichens, fresh water & marine microbial aggregates
- development of innovative tools for the analysis, integration and visualization of comprehensive „omics“ datasets
Subgroup „Pathogenomics“ (Dr. Jan Pané-Farré)
Dr. Jan Pané-Farré
The function of the SigB-regulon in Staphylococcus aureus. Many of the genes controlled by the alternative sigma-factor SigB in S. aureus encode for proteins of unknown function. We use a multi-disciplinary approach combining different omics, genetic, molecular biology and fluorescence microscopy techniques to study the function of selected members from this group of proteins. In addition, we are also interested in how the activity of SigB is regulated and under which conditions the expression of the SigB-regulon is advantageous for the cell.
Subgroup „Data analysis and Data integration“ (Dr. Stephan Fuchs)
Dr. Stephan Fuchs
Very large dataset are generated particularly by omics technologies and meta studies. We are working on global analysis strategies and how to integrate data from different omics technologies such as Transcriptomics, Proteomics, and Metabolomics. By this new insights can be provided into the adaptive physiology and pathogenicity of bacteria (e.g. Staphylococcus aureus). Furthermore, we are interested in complex microbial communities (microbiomes) and are working on appropiate meta analysis workflows. Prophane (http://www.prophane.de) and Aureolib (http://aureolib.de) are examples of successful data analysis and, respectively, integration ressources initiated by our group.
Industrial Proteomics“ (Prof. M. Hecker(retired), Dr. B. Voigt, Prof. K. Riedel)
- Analysis of the physiology and stress responses of bacteria used in industrial fermentations such as Bacillus licheniformis, B. pumilus and Ralstonia eutropha,with gel-based and gel-free proteomics (close collaboration with partners from industry such as the Henkel KGaA, BASF and AB Enzymes GmbH)
- High throughput proteomics for analysing samples from industrial fermentation we receive from our partner companies
- Modification of proteins during growth of bacteria (phosphoproteome, acetylome)
- Modifications of secreted proteins, studies of the exoproteome of bacteria
- Analysis of marine bacteria using proteomics techniques
- Proteomics service for academic and industrial partners.
Further Reading & Key Publications:
- B. Voigt, C.X. Hieu, K. Hempel, D. Becher, R. Schlüter, H. Teeling, F.O. Glöckner, R. Amann, M. Hecker, T. Schweder (2012) Cell surface proteome of the marine planctomycete Rhodopirellula baltica. Proteomics 12, 1781-1791
- M. Kleiner, C. Wentrup, C. Lott, H. Teeling, S. Wetzel, J. Young, Y.-J. Chang, M. Shah, N.C. VerBerkmoes, J. Zarzycki, G. Fuchs, S. Markert, K. Hempel, B. Voigt, D. Becher, M. Liebeke, M. Lalk, D. Albrecht, M. Hecker, T. Schweder, N. Dubilier (2012) Metaproteomics of a gutless marine worm and its symbiotic microbial community reveal unusual pathways for carbon and energy use. Proc Natl Acad Sci U S A. 109, E1173-1182
- C. Kaddor, B. Voigt, M. Hecker, A. Steinbüchel (2012) Impact of the core components of the phosphoenolpyruvate-carbohydrate phosphotransferase system, HPr and EI, on differential protein expression in Ralstonia eutropha H16. J Proteome Res. 6, 3624-3636
- P. Golec, J. Karczewska-Golec, B. Voigt, D. Albrecht, T. Schweder, M. Hecker, G. Węgrzyn, M. Łoś (2012) Proteomic profiles and kinetics of development of bacteriophage T4 1 and its rI and rIII mutants in slowly growing Escherichia coli. J. Gen. Virol. 94, 896-905
- H. Celik, J.C. Blouzard, B. Voigt, D. Becher, V. Trotter, H.P. Fierobe, C. Tardif, S. Pagès, P. de Philip (2013) A Two-Component System (XydS/R) Controls the Expression of Genes Encoding CBM6-Containing Proteins in Response to Straw in Clostridium cellulolyticum. PLoS One, 8, e56063
- B. Voigt, R. Schroeter, B. Jürgen, D. Albrecht, S. Evers, J. Bongaerts, K.-H. Maurer, T. Schweder, M. Hecker (2013) The response of Bacillus licheniformis to heat and ethanol stress and the role of the SigB regulon. Proteomics 13, 2140-2161
- R. Schroeter, T. Hoffmann, B. Voigt, H. Meyer, M. Bleisteiner, J. Muntel, B. Jürgen, D. Albrecht, D. Becher, M. Lalk, S. Evers, J. Bongaerts, K.-H. Maurer, H. Putzer, M. Hecker, T. Schweder, E. Bremer (2013) Stress responses of the industrial workhorse Bacillus licheniformis to osmotic challenges. PLoS One 8, e80956
- S. Handtke, R. Schroeter, B. Jürgen, K. Methling, R. Schlüter, D. Albrecht, S. van Hijum, J. Bongaerts, K.-H. Maurer, M. Lalk, T. Schweder, M. Hecker, B. Voigt (2014) Bacillus pumilus reveals a remarkably high resistance to hydrogen peroxide provoked oxidative stress. PLoS One 9, e85625
- M. Raberg, B. Voigt, M. Hecker, A. Steinbüchel (2014) A closer look on the polyhydroxybutyrate- (PHB-) negative phenotype of Ralstonia eutropha PHB-4. PLoS One. 9, e95907
- U. Brandt, C. Waletzko, B. Voigt, M. Hecker, A. Steinbüchel (2014) Mercaptosuccinate metabolism in Variovorax paradoxus strain B4-a proteomic approach. Appl Microbiol Biotechnol. 98, 6039-6050
Technical and methodological development for state-of-the-art proteomics
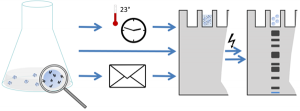
Sample processing is crucial for sensitive and comprehensive proteomic analyses, especially prior to mass spectrometric analysis in LC-MS/MS systems. Here we have developed and adapted a wide range of sophisticated protocols. Their elaborate use enables us to characterize proteomes of different microbes, both for our independent research and in cooperation with numerous national and international partners.
Some examples are:
Purification and enrichment of extremely diluted proteome samples
We have succeeded in establishing a protocol for enrichment of highly dilute samples, including proteins found in body fluids and excreted into growth media. It is based on commercially available affinity beads that have to be conditioned for this purpose in a proper fashion. Additionally most sensitive purification of proteome samples is possible, e.g. to clean up samples from salts or nucleic acids, that would affect following mass spectrometrical measurement. As a striking feature, this purification protocol further allows for shipping/ sharing proteome samples without elaborate cooling logistics.
Bonn et al. PMID: 24987932.
Establishment and optimization of new methods
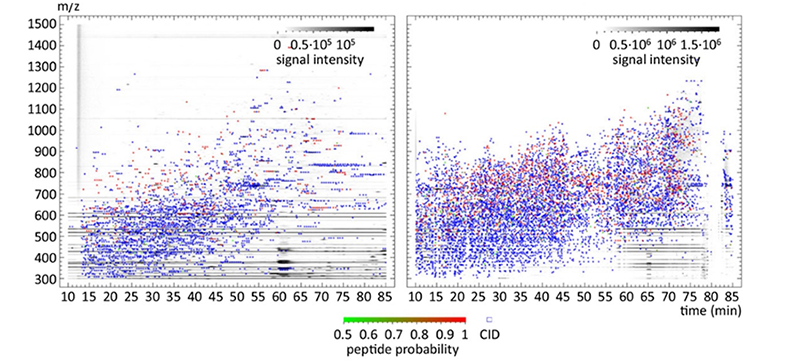
For mass spectrometry based proteomics we rely mainly on three technical fields that represent the modern, discovery driven, quantitative proteomics : Shot-gun proteomics/ data dependent mass spectrometry, SRM-based/ targeted mass spectrometry and data independent mass spectrometry. To achieve maximum performance we are strongly dependent on technological advances by mass spectrometric equipment. But this high-end-equipment must be also exploited in a proper way, workflows have to be optimized or newly developed. These efforts are not less important as frequently protocols decide over scientific output rather instrumentation.
Application of proteomics in biological and biotechnological research
Our research opens a broad variety of applications in several biological fields, thus applied proteomics could be addressed for different scientific questions in general, physiological, environmental or infection related microbiology. Therefore our workflows have been widely used in multiple studies, some examples are:
Biology of infection and stress physiology
The fast adaptation of pathogenic microbes to host defenses or new antimicrobial therapies makes it necessary to develop novel therapeutic strategies to combat such infections. Pre-requisite is a better understanding of these processes of bacterial adaptation which are often anchored deeply in fundamental biology (evolutionary mechanisms, horizontal gene transfer). As comparative proteome expression analyses give qualitatively improved insights into these adaptational networks, we are involved in numerous physiological studies. Some examples are:
- Maaβ et al. PMID: 24878497.
- Huja et al. PMID: 25118243.
- Wenzel et al. PMID: 24706874.
- Hoffmann et al. PMID: 24373249.
- Pribyl et al. PMID: 24387739.
- Lappann et al. PMID: 23893116.
Absolute protein quantification in systems biology
In systems biology, the generation of interdependent data requires the integration of large scale ‘omics’ studies each yielding absolute data for different levels of cellular components like DNA, RNA, proteins or metabolites. Although each single ‘omics’ level data leads to unique information, the analysis of the proteome is particularly valuable due to the nature of proteins as the real effectors of cellular metabolism (enzymes and structural determinants) throughout all kingdoms of life. For generation of absolute data in systems biology approaches we rely on three different workflows:
1. Determination of absolute quantities of single proteins in complex mixtures by AQUA and targeted proteomics
2. 2D Gel based absolute quantification combining AQUA and state-of-the-art 2D-PAGE
3. Absolute quantification based on LC-MS techniques (TOP3 on data independent MS- platforms)
- Muntel et al. PMID: 24696501.
- Maaβ et al. PMID: 24878497.
- Buescher et al. PMID: 22383848.
Environmental Metaproteomics
Elucidation of processes in nature has driven scientists ever since – and has gained even more attention in recent times with new technologies at hand allowing for both broad and detailed analyses of environment. Fascinating processes taking place e.g. in coastal areas, soil, microbial mats or in the human gut system that could not or not sufficiently be analyzed with technical opportunities of the pre-proteomics era. For extremely complex proteomic samples and resulting data, it is crucial to rely on state-of-the-art mass spectrometry methods as well as on highly sophisticated software capable of dealing with the information attained.
Within a network of collaborating Northern German Scientists we have succeeded in getting a glimpse into the complex bacterial successions in the North Sea at Heligoland based on multiple ‘omics techniques including proteome data.
- Xing et al. PMID: 25478683.
- Teeling et al. PMID: 22556258.
Research Group under construction 1
Research Group under construction 2
Department „Microbial Ecology“ (Prof. C. Gliesche)
The AG Microbial Ecology investigates the impact, diversity and function of microorganisms playing a crucial role in global biogeochemical cycles. Special focus lies on microbial communities in coastal sediments of the southern baltic sea. Molecular-biological as well as sensor-based technologies are employed to provide new insights into microbial processes at border surfaces and in microhabitats.
Senior Research Group „Stress Physiology I“ (Prof. M. Hecker (retired))
Senior Research Group „Stress Physiology I“ (Prof. M. Hecker (retired))
- The Bacillus subtilis σB-dependent general stress response: More than 200 genes belong to the σB-dependent general stress regulon. These proteins equip the non-growing cells with a multiple, non-specific and preventive stress resistance in anticipation of “future stress”. The “fine-tuned” gene regulation and function of single stress proteins in the establishment of a global resistance against heat, ethanol, oxidative and osmotic stress are currently analyzed. Furthermore, the integration of the general stress regulon into a highly sophisticated adaptational network is being investigated.
- “The Bacillus subtilis Clp machinery”: Regulation of the stress-inducible clp genes depends predominantly on the transcriptional CtsR repressor. CtsR is targeted by the ClpEP and ClpCP proteases during heat stress. Moreover, ATP-dependent proteolysis mediated by Clp proteases can also be observed during general stationary-phase phenomena, such as glucose starvation, competence development and sporulation. Mechanisms that finally result in the inactivation and degradation of transcriptional regulators and proteins, which lost their “duty” are currently investigated. Moreover, the role of the McsB arginine kinase, an adaptor protein of the ClpCP protease, is in the focus of our research.
Further Reading & Key Publications:
- Reder et al. Cross-talk between the general stress response and sporulation initiation in Bacillus subtilis – the σB promoter of spo0E represents an AND-gate. Environ Microbiol (2012) 14(10), 2741–2756. PMID: 22524514.
- Reder et al. The modulator of the general stress response, MgsR, of Bacillus subtilis is subject to multiple and complex control mechanisms. Environ Microbiol (2012) 14(10), 2838–2850.PMID: 22812682.
- Elsholz et al. CtsR, the Gram-positive master regulator of protein quality control, feels the heat. EMBO J. 2010 Nov 3;29(21):3621-9.
- Elsholz et al. Global impact of protein arginine phosphorylation on the physiology of Bacillus subtilis. PNAS 2012 May 8;109(19):7451-6 PMID: 22517742.
Senior Research Group „Applied Microbiology“ (Prof. F. Schauer (retired))
The analysis of the chemical structure of metabolites from bacteria, yeasts and filamentous fungi by LC-MS, GC-MS and NMR leads to the description of new degradation pathways of oil components (alkanes, cycloalkanes, alkylbenzenes), biarylic components (biphenyls, dibenzofurans, diphenylethers, bisphenols), herbicides, disinfectants or special pharmaceuticals and drugs. The isolation of a great variety of microorganisms results in the establishment of a strain collection SBUG of 6.500 biotechnological applicable microorganisms.
