Main research areas
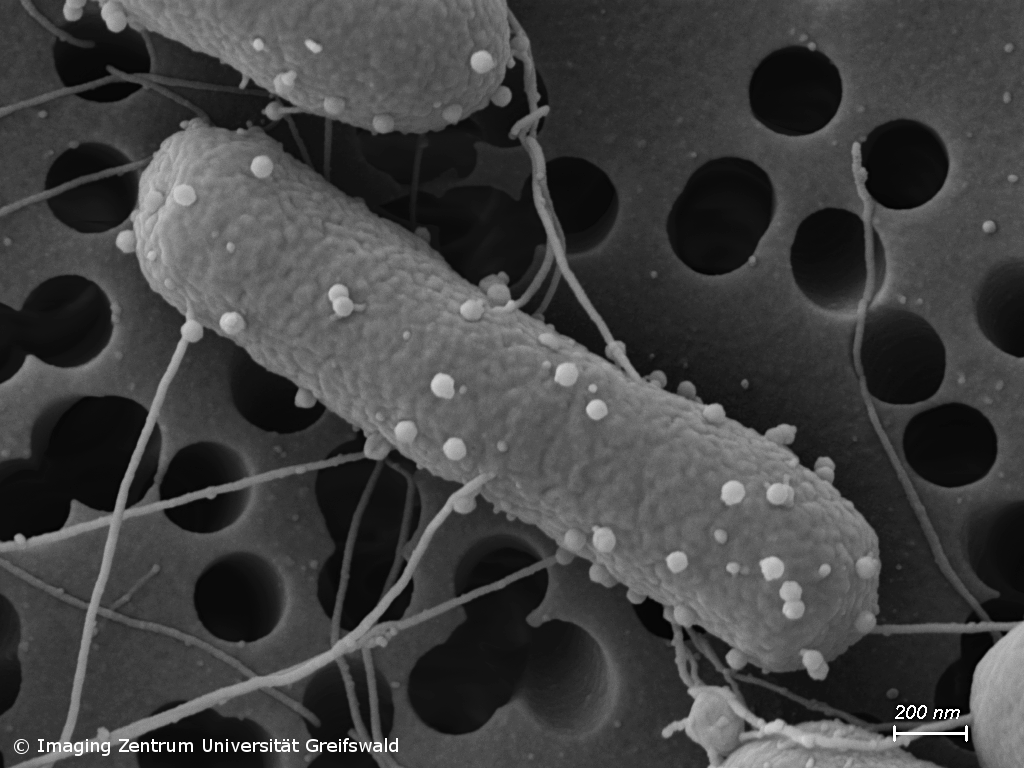
Antimicrobial resistance (AMR) and novel antimicrobial strategies
Infections with antimicrobial resistant pathogens are one of the major threats of mankind. Already today, clinicians have to deal with bacteria that show resistance to all currently available antibiotics. Our group studies occurrence and transmission of pathogens carrying AMR genes, e.g. in waste water and soil, but we also conduct research on the development of alternative antimicrobial therapies. For instance, we analyze the activity and mode of action of novel synthetic and natural substances against problematic bacterial pathogens, but also hunt for bacteriophages with the potential to be further developed to be perspectively used in phage therapy.
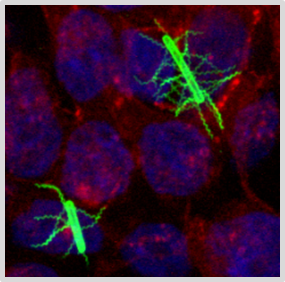
Pathophysiology of Clostridioides difficile (CDinfect)
The strictly anaerobic bacterium C. difficile is one of the most problematic pathogens causing nosocomial and relapsing infections. The intestinal pathogen can survive in the gut and establish infections due to its ability to form highly resistant endospores and to withstand the harsh conditions in the human intestinal tract. Despite the importance of C. difficile infections, the physiology of the pathogen is still not very well understood. We aim to pinpoint the main metabolic pathways and physiological efforts that empower C. difficile to thrive in the intestinal tract and to cause severe infections in humans.
Membrane structure and membrane-active compounds
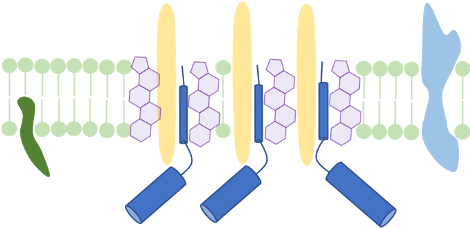
The cellular membrane separates the intracellular space from the exterior. It does not only accounts for the cell’s integrity, but also enables transport-, signal reception- and transduction processes. We make use of membrane proteomics and microscopic techniques to investigate the impact antimicrobial substances might have on membrane structure and function. Currently, we analyze changes of functional membrane microdomains in C. difficile in the presence of infection-relevant bile acids. We propose that host- or microbiome-derived membrane-active substances could have a combinatory effect with antimicrobial substances that are therapeutically used in infection.
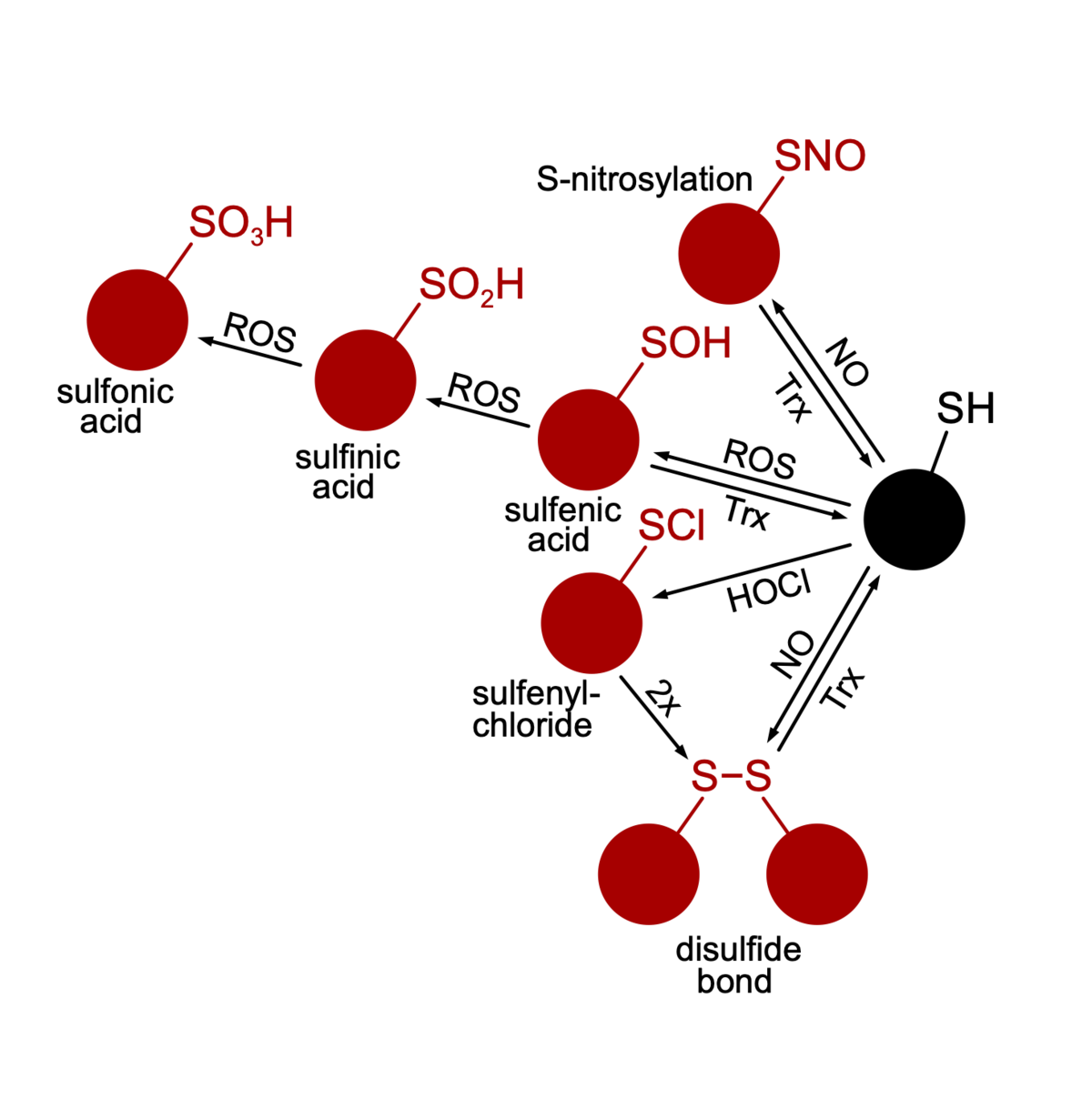
Oxidative stress and anti-oxidative strategies (BiOx GRK1947, FlavOx DFG)
Life is based on redox reactions, i. e. the cell is full of molecules that reduce or oxidize each other. Some molecules have a bigger oxidative potential than others. Especially in the presence of oxygen, reactive oxygen species (ROS) evolve that have the potential to oxidize cellular structures instigating signal transduction pathways or damaging macromolecules. Microorganisms that depend on oxygen in their metabolism feature sophisticated stress response-, repair- and ROS-detoxification mechanisms. One could assume that strictly anaerobic organisms are devoid of such mechanisms since they grow in habitats free of oxygen. However, many strictly anaerobic bacteria are capable of an oxidative stress response and can thus survive short periods of oxygen and ROS presence. With respect to anaerobic pathogens, the ability to survive periods of oxidative stress can decide on whether the pathogen can assert in the host or not. We study the oxidative stress response in C. difficile and thereby elucidate ROS sensors, signal transduction pathways, protective molecules and repair mechanisms. A special focus is put on oxidized proteins. Using mass spectrometry-based proteomics and redox-proteomics we aim to pinpoint redox switches and mostly vulnerable structures in the C. difficile cell.
Soil microbiome and its impact on agricultural crops (AgriBiom T!Raum One Health BMBF)

There is virtually no space on earth that is devoid of microbes. Specific niches feature their own specific microbial community – a niche-specific microbiome that plays an often still unknown role for the stability and equilibrium of this niche.Our expertise in mass-spectrometry-based proteomics and meta-proteomics enables us to perform in-depth analyses of the soil microbiome of agricultural grounds. On the one hand our aim is to unravel the microbes and their metabolic capabilities that are beneficial for the growth and health of agricultural crops and on the other hand identify microbial parameters that indicate deleterious effects emanated from the soil microbiome.
Research projects
Current projects
- T!Raum One Health
AgriBiom "Establishment of a standard diagnostic test for the microbiome of agricultural soils"
Mai 2026 - Exploring the ecology and evolutionary of phages to unveil their potential utility for humans (UtiliPhage)
März 2027
Completed projects
- BIOx (FKZ GRK 1947): Biochemical, biophysical effects of reactive oxygen and nitrogen species on biological membranes,
TP C4: The membrane of the intestinal pathogen C. difficile as the first line of defense during infection-related oxygen and bile acids challenge - FlavOx (FKZ SI 1678/3-1): Elucidation of the role of flavodoxins in intestinal pathogens Clostridioides difficile
- Molecular basis for host adaptation and pathogenicity of zoonotic hantaviruses (HANTadapt)